OutbreakTrees: an Open-access database of infectious disease transmission trees
Transmission trees describe who infected whom during outbreaks of infectious diseases (see example tree below). These data are routinely collected through resource intensive methods including case finding, contact tracing, detailed epidemiological interviews, and probabilistic reconstruction. Statistics describing transmission trees, including the mean and variation in the number of infections each infected individual causes, reveal aspects of pathogen nature that govern disease spread and effective intervention strategies.
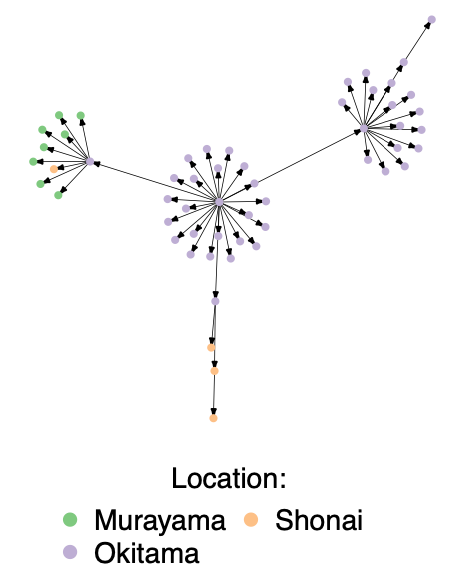
We compiled and standardized over 350 transmission trees from published literature representing 16 directly transmitted infectious diseases in an effort to make transmission tree data more accessible to the research community. Trees are downloadable as igraph objects, where nodes represent individuals, edges represent person-to-person transmission, and node attributes include valuable information about infected individuals described in the publication (e.g., location, occupation, relationship to infector). In addition, the database reports summary statistics about each tree (e.g., outbreak size), metadata (e.g., country, year, any assumptions we made in entering), and a link to the original source. Those interested are encouraged to visit our new portal (outbreaktrees.ecology.uga.edu) to discover, visualize, and download data. For a thorough description of inclusion criteria, see the paper: Taube, Miller, and Drake 2021.
In the paper, we also demonstrate the utility of this database by exploring key questions about superspreading including the relationship between superspreading and outbreak size, the timing of superspreading, and whether superspreaders tend to infect other superspreaders.